Introduction
This web-based tool developed at RAS Initiative at Frederick National Lab for Cancer Research, namely RAS Pathway DepMap Correlation Network Explorer, is intended to help study and infer the potential context-dependent gene-gene relations on top of correlation networks derived from DepMap genome-wide CRISPR screening data (DepMap Portal), which was originally created by DepMap Consortium (DMC) and hosted by BROAD Institute, and primarily focus on the context of RAS signaling with derived context-dependent functional modules, GPMTC (Good Paired Mutual Top Correlated) genes, as well as gene-gene correlation relations.
Below we briefly introduce the main features of the tool, which visualizes, manipulates, and analyzes the RAS-pathway focused and genome-wide ready correlation gene networks embedded with functional modules and GPMTC genes.
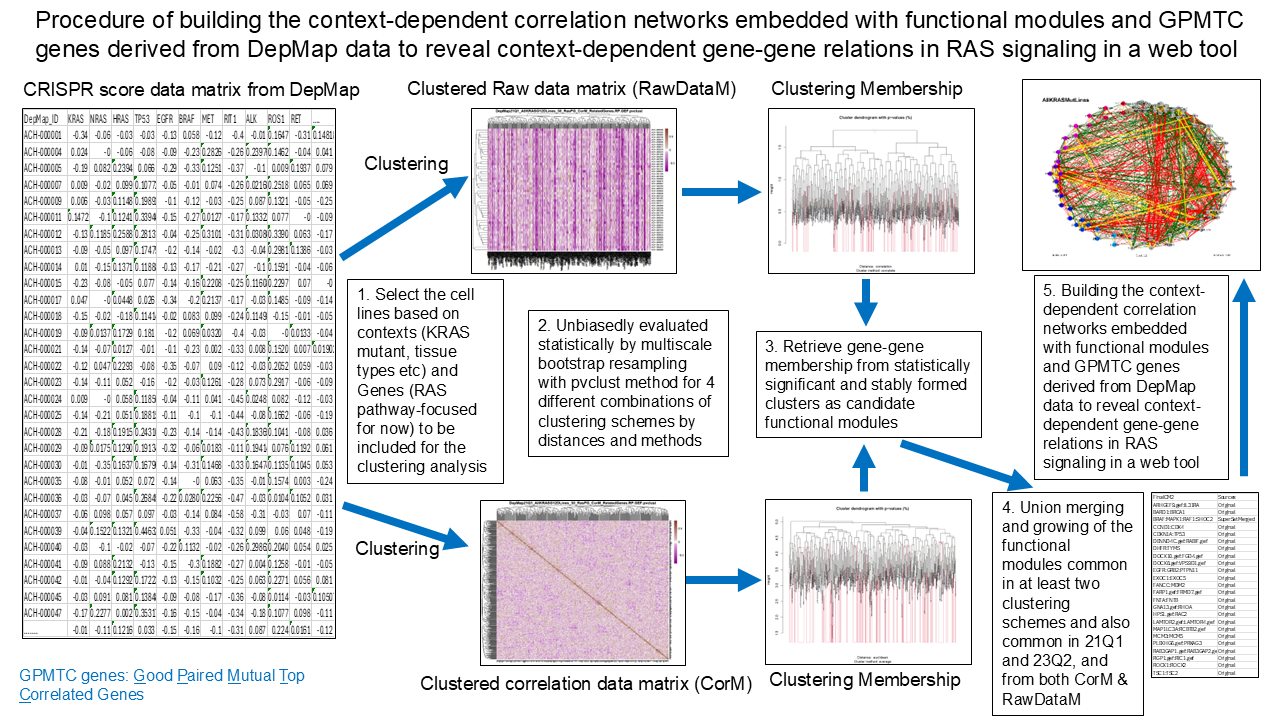
Functional modules were derived by assessing the uncertainty in hierarchical cluster analysis using the dependency data in raw data matrix as well as in the correlation matrix unbiasedly and statistically by multiscale bootstrap resampling with pvclust method for 4 different combinations of clustering schemes by distances and methods. The 21Q1 and 23Q2 versions of DepMap data were used to test the robustness of the revealed functional modules. In the "circular" layout, member genes in each functional module are displayed in each small circular area for the corresponding module in an assigned rainbow color, although other genes such as RAS genes or closely related genes commonly critical in RAS pathway are also included, if not in derived function modules,as colored in gray nodes, which are also organized into small circular area in "circular" layout. All the genes are arranged along the large peripheral circle of the whole network in "circular" layout although other layout styles can be used. The GPMTC genes, which are mutual top correlated genes for each other in each pair of GPMTC genes, are connected with gold-colored lines, with many of the GPMTC genes also as genes of functional modules. Those not belonging to functional modules are displayed as gray-colored nodes.
The lines connecting genes represent correlation levels by thickness of the lines: green for positive and red for negative correlation. Only significant correlation levels will be visible as lines displayed in the network. As mentioned, gold lines connect the GPMTC genes. The default layout of the network is "circular," although you can change to other layout styles with "Layout" options for the same set of genes in the current displayed network.
You can choose to display the default network for a chosen cell type (gene mutation status, e.g., RAS genes, BRAF; tissue types, e.g., Pancreas, Lung, Colon, etc.; RAS dependency status) by selecting from a list of different input cell types/contexts. You can also visualize the relations of the same set of genes in another cell type by choosing a new cell type to be changed on the current network.
The font size of genes in the network and the title of the network can also be changed. You can manipulate the network in many different ways: add your interested genes, if not present, to the currently displayed or default network (note: if genes are not in DepMap datasets, they may not be added, although 17k genes are covered); visualize a subset of the current network; find and add top bridge gene(s) (designated as the top dual correlated gene(s) with the top product(s) of correlation coefficients for both seed genes as the potential bridge gene(s)) for each pair of interested seed genes; delete the gene(s) from the current network; expand the current network by functional modules, GPMTC genes by default, or by top correlated genes (up to top 100, can be increased to more if needed) as defined by you; and undo the previous network manipulation.
You can save the current network into a designated .csv file, which can be used to re-create the network later, as long as you use the same cell type/context. You can also save the image of the current network into an image file for a chosen format.
This tool is dynamic, and you are welcome to approach us (RAS Initiative Bioinformatics Research Team) to potentially get better cell types or contexts that meet your needs. If your interested cell type or context is not available in the current dropdown list of cell types/contexts, you can either choose a closely related cell type in the list or ask if we can create that cell type specifically for your purpose, as long as the DepMap data has enough numbers of the cell lines for that cell type/context. Please first check out the last section at the bottom of this web page as below for more details. Also, if possible, we will create the cell type and context and include it in the list of selectable cell types for customized usage.
Below in the next section, we included a few tutorial videos to help you get started.
In addition, please check out the last section entitled "Useful Information about the Details and Numbers of Cell Lines for Each Cell Type/Context Currently Available in the Tool" at the bottom of this webpage, which not only lists the cell types/contexts available in the tool and related details but also informs whether a corresponding cell type/context has sufficient numbers of cell lines for deriving the gene-gene correlation, function modules and GPMTC genes, since generally the more cell lines available for the cell type/context, the inferred gene-gene relations would be more reliable.
To access the tool, use the following hyperlink: RAS Pathway DepMap Correlation Network Explorer.